Structural biology of RNA-protein complexes
PI: Carine Tisné
Microbial Gene Expression Laboratory
2-year contract
The research of the team “RNA biogenesis, architecture and interactions” led by C. Tisné is focused on structural biology of RNA-protein complexes involved in RNA maturation. RNA is heavily co- or post-transcriptionally modified, with the largest diversity of modifications found in tRNAs. The recent discovery of modifications in mRNAs and their involvement in the regulation of gene expression has fueled a new field of investigation in this oldest area of RNA research, which has so far focused mainly on tRNAs and rRNAs. Given the widespread presence of post-transcriptional RNA modifications in all classes of RNAs and in all areas of life and their dynamics, it is clear that RNA processing pathways and RNA functions must have evolved under the influence of RNA modifications. The interplay between RNA processing and RNA modifications has received little attention so far. We want to understand how RNA modifications and processing together determine RNA maturation and, in turn, control or influence RNA functions. Applicants for this position should propose a project using structural biology to address these questions of RNA biology.
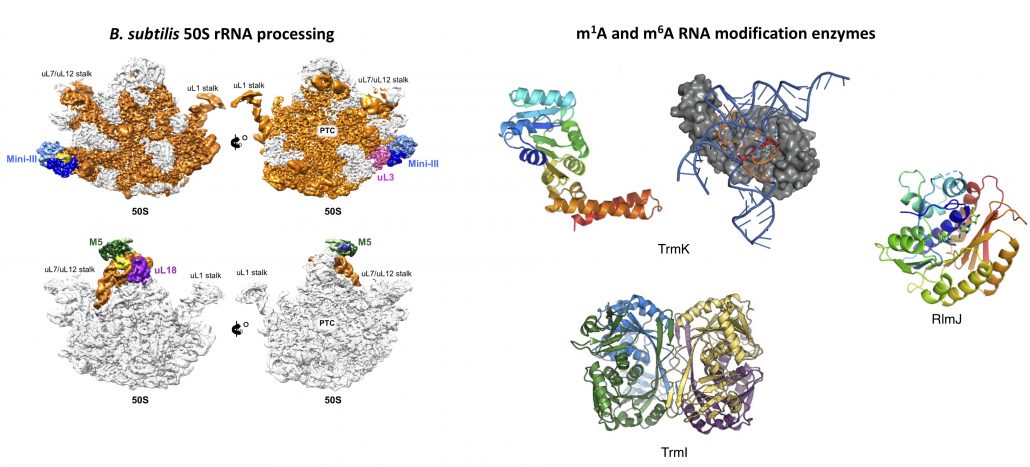
Oerum S. et al (2021), A comprehensive review of m6A/m6Am RNA methyltransferase structures. Nucleic Acids Res. 49(13):7239-7255. doi: 10.1093/nar/gkab378.
Oerum S, Dendooven T, Catala M, Gilet L, Dégut C, Trinquier A, Bourguet M, Barraud P, Cianferani S, Luisi BF, Condon C, Tisné C. (2020), Structures of B. subtilis Maturation RNases Captured on 50S Ribosome with Pre-rRNAs. Mol Cell. 80(2):227-236. doi: 10.1016/j.molcel.2020.09.008.
Barraud P, Gato A, Heiss M, Catala M, Kellner S, Tisné C. (2019), Time-resolved NMR monitoring of tRNA maturation. Nat Commun. 10(1):3373. doi: 10.1038/s41467-019-11356-w.
For more information on host lab, click here:
http://www.ibpc.fr/UMR8261/AccueilEN/Theme%20CT/ThemeCT.html

Carine Tisné’s team

PI Carine Tisné