Technical platforms
Functional Genomics Plateform
Bioformatics plateform
Recently set up at IBPC, the functional genomics platform puts its know-how at the service of the IBPC units and collaborates with the scientific community.
The functional genomics platform is also equipped with several apparatuses for gel/image processing and analysis, notably a Typhoon FLA9500 (GE), 2 Gel Docs (Biorad) and developer for autoradiograms (Curix), as well as a scintillation counter (Hidex) for processing radioactive samples
Contact: bioinfo@ibpc.fr
Scientific supervisor : Alexandre Maes (alexandre.maes@ibpc.fr)
Objectives
- Advise and train IBPC researchers for routine bioinformatics analyzes.
- Develop new analysis and visualization tools adapted to the latest technologies and users.
- Develop collaborative research projects with the different IBPC teams and research units.
- To bring together the institute’s bioinformaticians and pool the many skills and resources in biological data analysis.
Thematic specificities and expertise
- NGS (genomic and transcriptomic), proteomic and quantitative proteomic analyses.
- Sequencing, assembly and annotation of bacterial and eukaryotic genomes.
- Metagenomics and functional annotation.
- Evolutionary biology.
- Visualization and supervised exploration of large collections of biological data.
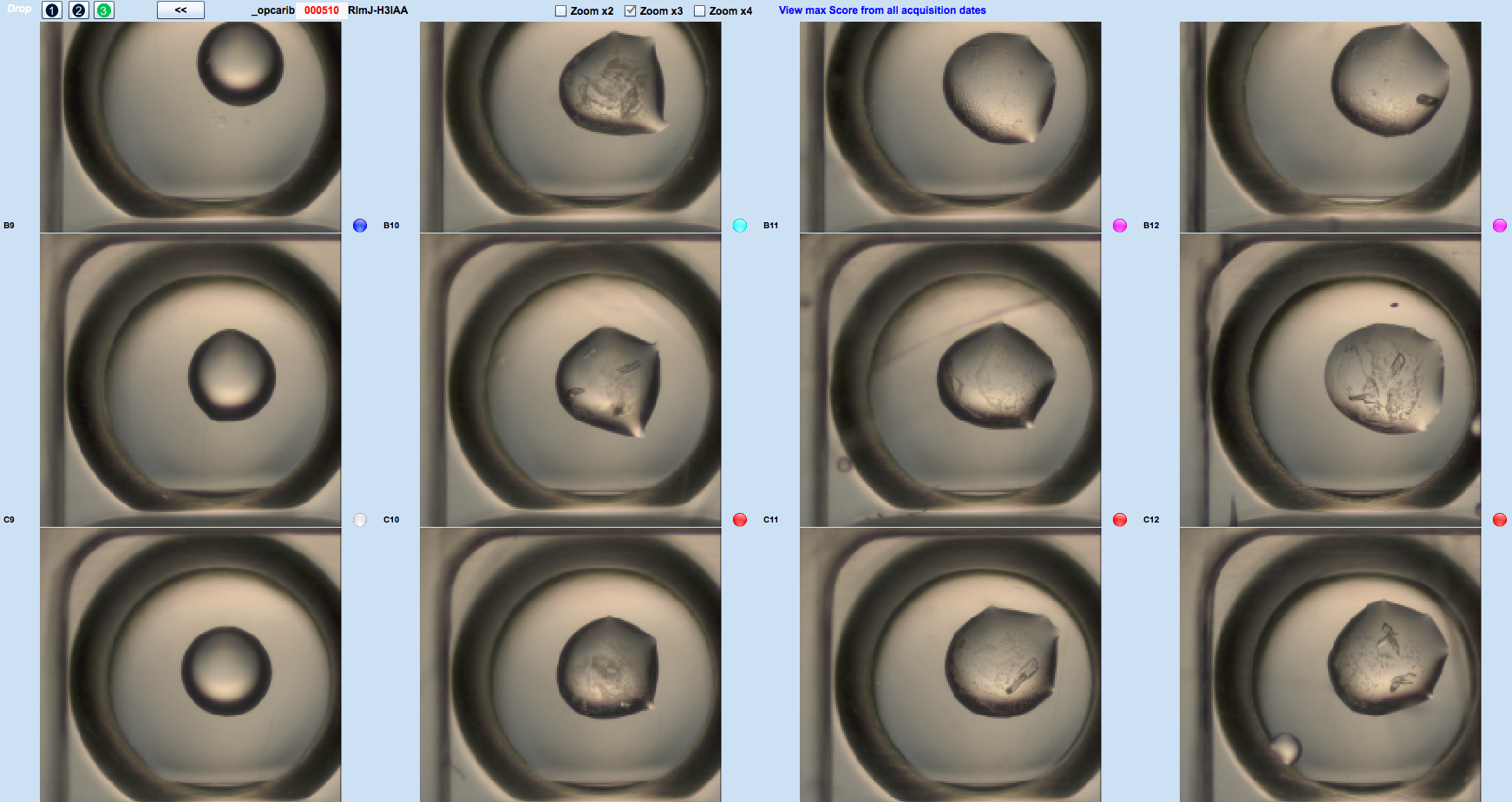
Crystallography
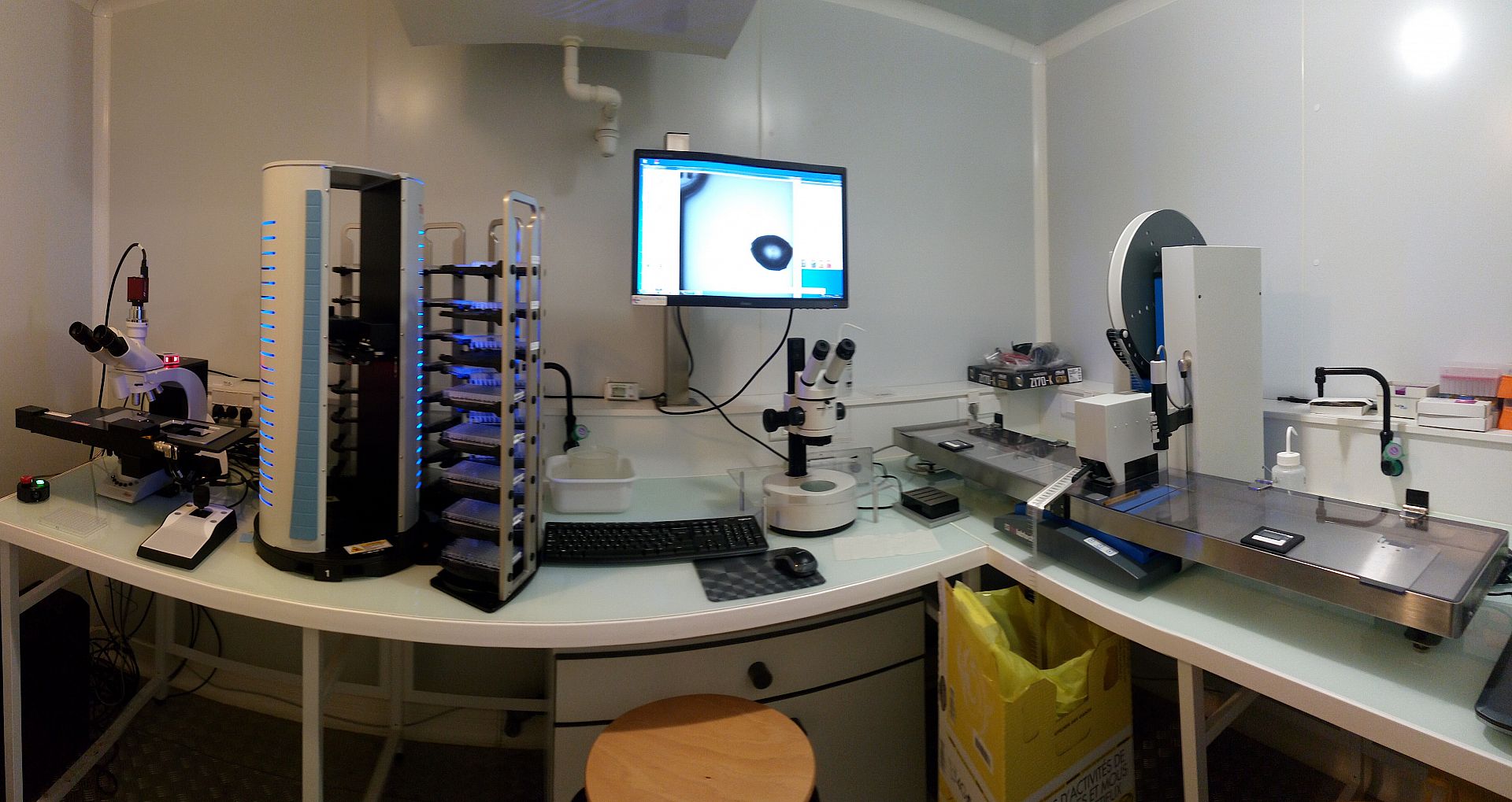
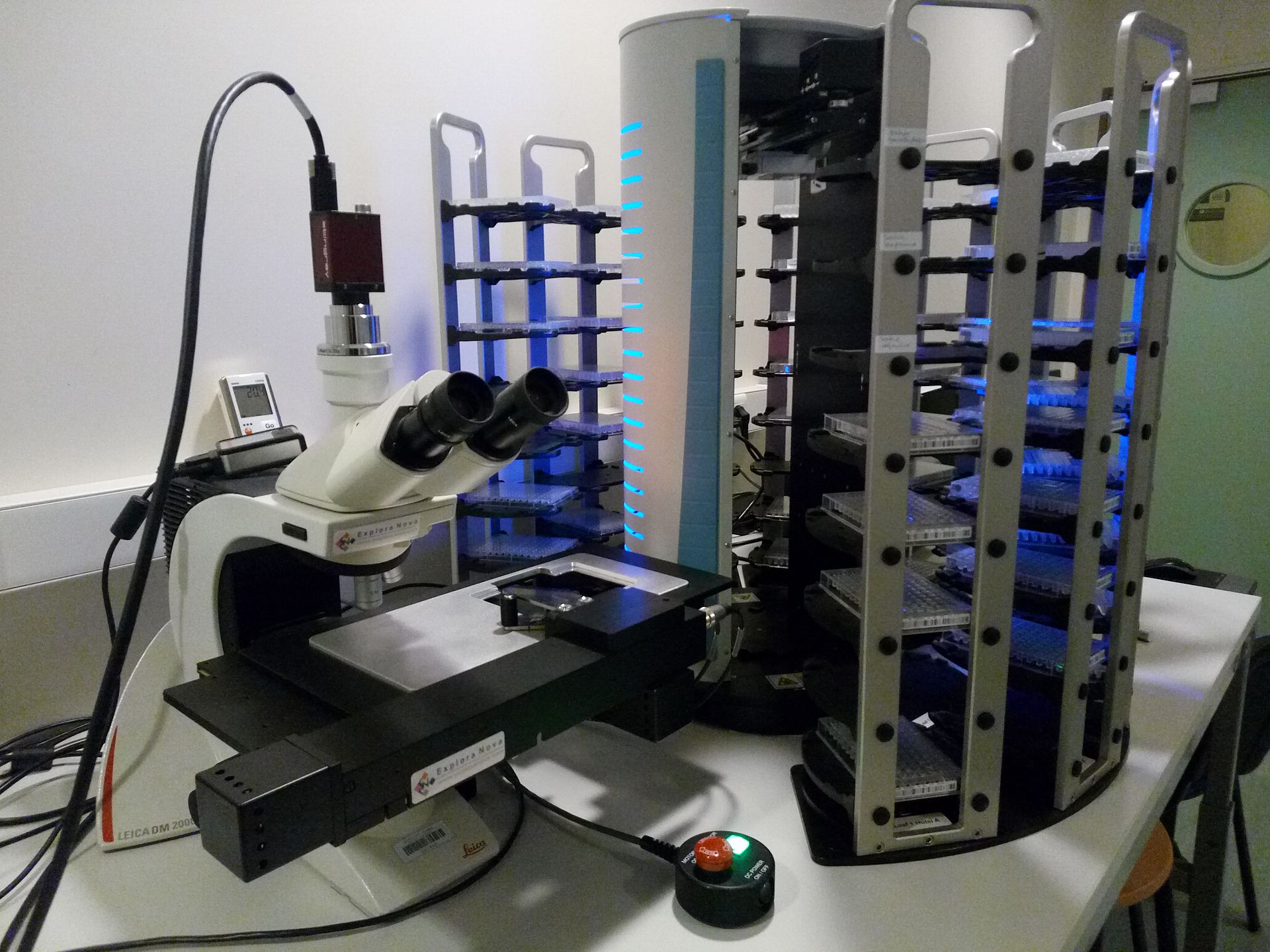
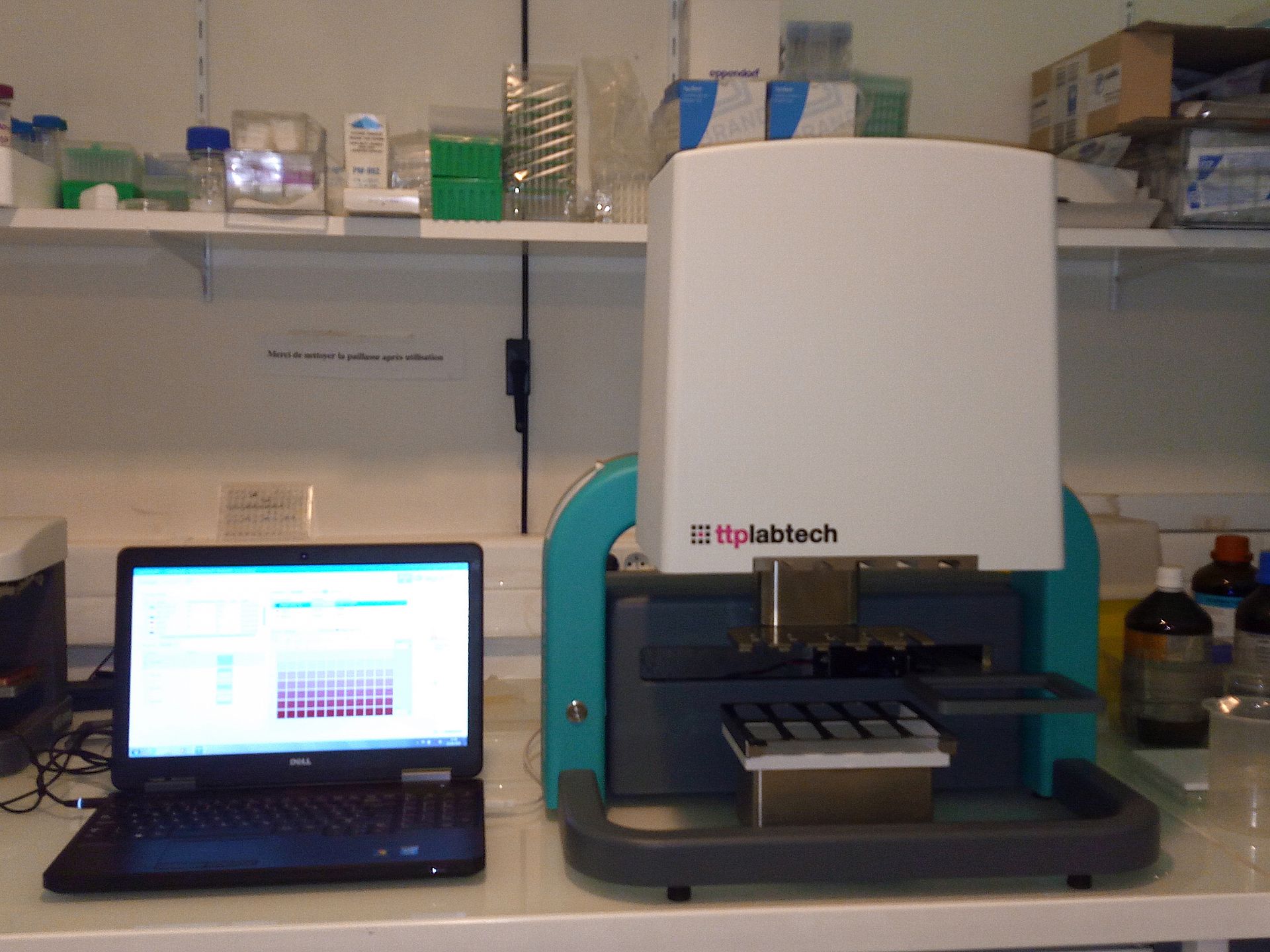
The Crystallography platform has evolved significantly since its inauguration on May 20, 2003 to meet to our current expectations. As of 2014 the platform is equipped with two latest-generation Mosquito pipetting robots allowing manipulation of biological macromolecules in viscous lipid phases and a distribution station (TTPlabtech) to prepare solutions used by the robots. The platform is equipped with incubators (18°C and 4°C), Zeiss microscopes and an automated system for visualizing crystals.
NMR
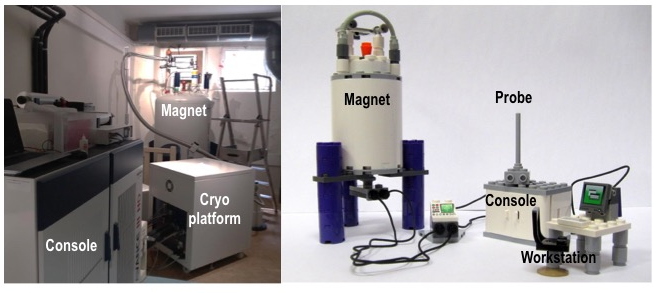
The IBPC’s biomolecular NMR platform is a platform of the research federation of the IBPC (FRC550). The platform serves the coordination of research in the field of nuclear magnetic resonance spectroscopy (NMR) within the IBPC. The platform provides the researchers, students and engineers of the IBPC with access to state-of-the-art infrastructure for the study of biological macromolecules and their complexes. The platform was installed in 2016 as part of the Equipex CACSICE. It consists of a Bruker Avance III-HD 700 MHz spectrometer and various liquid and solid probes. In 2018, thanks to additional funding from the Île-de-France region and the Labex Dynamo, the platform was equipped with a cryoplateform and a latest-generation liquid cryoprobe.
For autonomous users with adequate training in NMR spectroscopy, the NMR spectrometer time required for their projects is simply made available according to the availability of the machine. For users who do not have sufficient autonomy, expertise and support can be provided through collaborations with experienced users of the platform. Depending on the project, these expertise and support may concern all aspects of NMR studies with biological macromolecules in solution, including know-how in the production of isotopically labelled samples, the implementation of NMR experiments, data analysis and structural calculations. The platform is not able to provide services outside collaborations with experienced users of IBPC, and therefore invites the persons interested in NMR to discuss their projects with them

ACKNOWLEDGEMENTS
Sentence to include in any publication that uses results acquired at the IBPC biomolecular NMR platform :
« The authors acknowledge access to the biomolecular NMR platform of the IBPC that is supported by the CNRS, the Labex DYNAMO (ANR-11-LABX-0011), the Equipex CACSICE (ANR-11-EQPX-0008) and the Conseil Régional d’Île-de-France (SESAME grant) »
Proteomics
The Proteomics platform was installed at the IPBC in 2014 thanks to funding from LabEx DYNAMO and EquipEx CACSICE. Its equipment and know-how are available to the scientific community of the two consortia.
The objectives of the platform are :
To advise researchers on the techniques available in the platform based on their projects
To analyse samples from their preparation to the exploitation of the results
To develop new analytical techniques as a function of researchers’ needs (e.g. label free quantifiacation)
To train researchers in sample preparation, data analysis and exploitation
The Proteomics platform can function as a service or in collaboration mode as a function of researchers’ preferences.
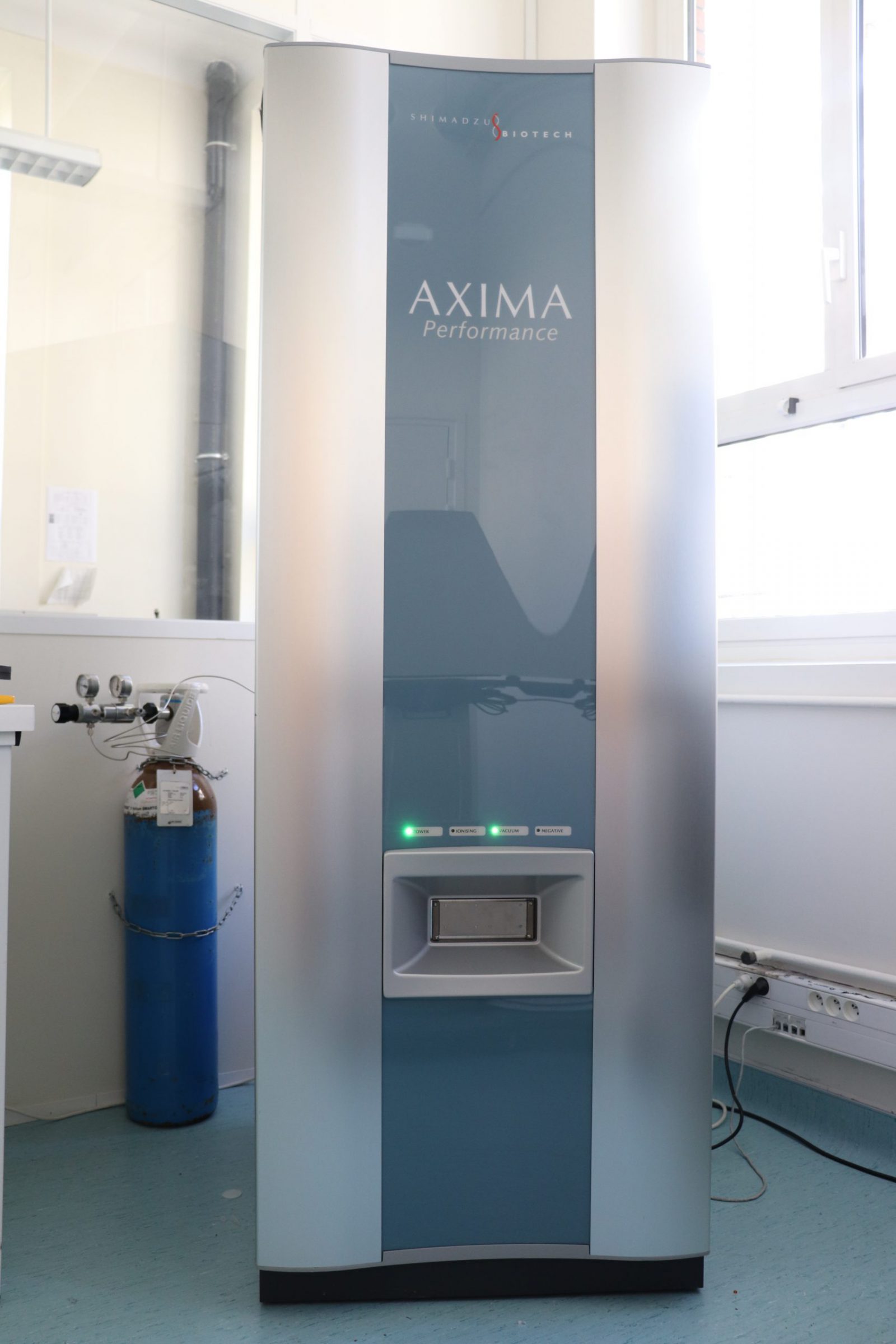
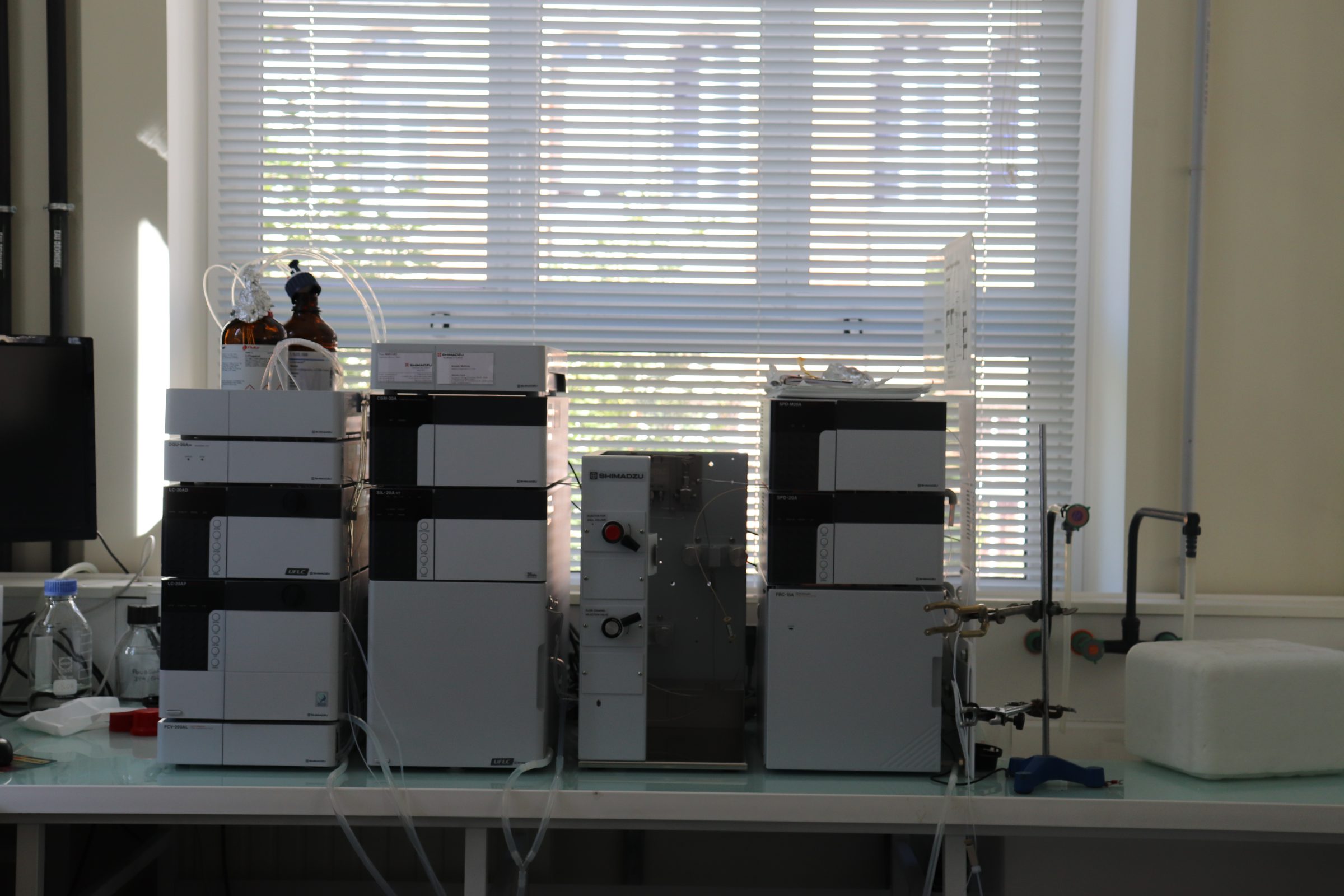
Equipment
The Proteomics platform is equipped with:
Analysis
The Proteomics platform proposes different types of services for sample preparation and analysis:
In-gel digestion of sample (spot or band)
Digestion of sample in solution
Purification of peptides (C18 column)
Identification of purified proteins (simple peptide mixture)
Identification of proteins in a complex mixture (following chromatographic separation)
Measurement of mass of intact proteins (in solution)
Quantitative proteomics (label free techniques)
Protocols
Protocols are provided, but contact with the Proteomics platform is recommended before any sample preparation is undertaken
A reservation schedule for the MALDI-TOF is available on line for trained personnel at the IBPC at https://resa-equipement.ibpc.fr/.
For all external requests for analysis or training, please contact hamon@ibpc.fr.
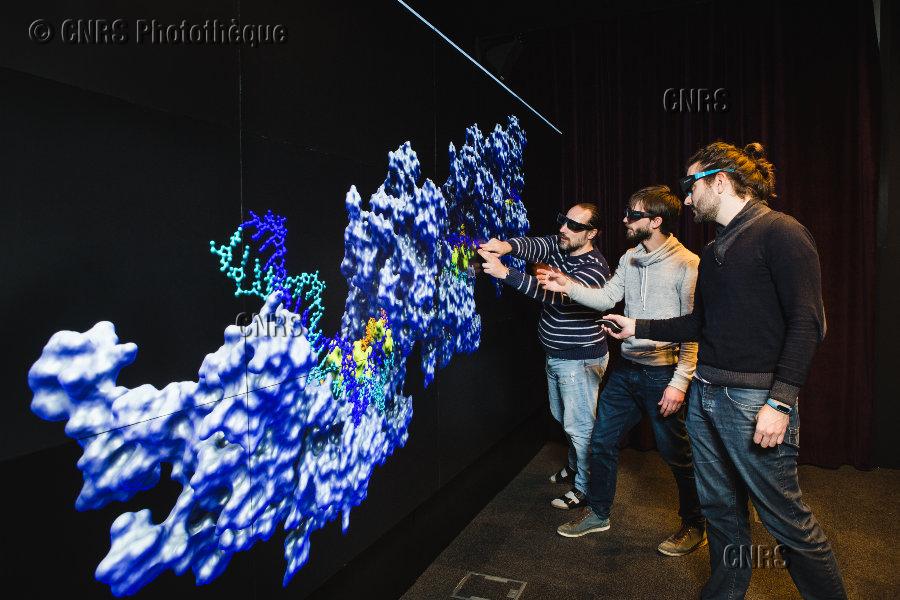
Visualisation wall
The visualization platform at the IBPC institute offers an 25 MPixel high-resolution high-dimension display wall with stereoscopic 3D representations for a semi-immersive experience allowing researchers to explore and manipulate their data.
The display wall consists of 12 stereoscopic displays arranged in a four column by three row matrix setup with a total dimensions of 4.4m wide by 1.8m high and a resolution of 7680 by 3240 pixels, addressed as a single screen through Windows, Mac OSX or Linux workstations.
It empowers researchers to visualize high resolutions images (from electron, confocal, etc microscopy) and visualize macromolecule structures and simulations trajectories in 3D.
EQUIPMENT
The platform is composed of :
– display wall with stereoscopic representation composed of 12 displays with a total dimensions of 4.4m wide by 1.8m high and a resolution of 7680 by 3240 pixels
– 3 workstations (Windows, Mac OS X, Linux) connected to the wall with 4 GPU cards in each.
– A control system (tablet, screen, keyboard and gyroscopic mouse) to interact with the wall.
SERVICE
The platform offers to researchers a high dimension and stereoscopic display to explore and manipulate their data. It allows to :
– Display high resolutions images from experimental procedure (Confocal Microscopy, Scanning Electron microscope, …)
– Display structure of macromolecules and trajectories of simulations in 3D (Pymol, Chimera, Coot, VMD, UnityMol, )
– Visualize simoutinlasy different type of data thanks to the high dimension.
– Display at the same time the content of 4 notebooks on the wall to facilitate information sharing during meetings.